| Organism | Source | Retrieve date | tRF Inclusion Criteria |
|---|---|---|---|
| Homo sapiens | tRFdb | 2022-11-12 | |
| tsRNA study | |||
| tsRBase | 2022-11-13 | function / expression experimentally validated | |
| tsRFun | 2022-11-12 | 1) Expressed in > 30% of the TCGA samples; or 2) have experimentally validated targets | |
| MINTbase v2.0 (04/04/2019) | 2022-11-15 | 1) [#datasets RPM>100]≥10 and [#datasets RPM>100]/[#datasets expressed]≥0.05, or 2) have experimentally validated targets in tRFTar |
|
| tRFexplorer (2019-05) | 2022-11-17 | fragment type: 5' Leader and Minimum RPM value: 10 | |
| tatDB | 2022-11-17 | MFE≤-20, Read count≥25, Unique hybrids≥1, Motif checked and Intro Excised unchecked | |
| OncotRF | 2022-11-17 | experimentally validated | |
| Mus musculus | tRFdb | 2022-07-16 | |
| tsRBase | 2022-08-21 | function / expression experimentally validated | |
| Caenorhabditis elegans | tRFdb | 2022-11-03 | |
| Danio rerio | tRFdb | 2022-11-01 | |
| Drosophila melanogaster | tRFdb | 2022-10-30 | |
| tsRBase | 2022-10-30 | function / expression experimentally validated | |
| Rattus norvegicus | tsRBase | 2022-11-10 | function / expression experimentally validated |
| Rhodobacter sphaeroides | tRFdb | 2022-11-06 | |
| Schizosaccharomyces pombe | tRFdb | 2022-11-09 | |
| Xenopus tropicalis | tRFdb | 2022-11-05 |
| Organism | Version | Retrieve date |
|---|---|---|
| Homo sapiens | GRCh38.p13 (GENCODE Release 42) | 2022-11-12 |
| Mus musculus | GRCm39 (GENCODE Release M31) | 2022-10-25 |
| Caenorhabditis elegans | WBcel235 (WormBase WS286) | 2022-11-03 |
| Danio rerio | GRCz11 (Ensembl release 108) | 2022-11-01 |
| Drosophila melanogaster | dmel_r6.32 (FlyBase FB2020_01) | 2022-10-25 |
| Rattus norvegicus | mRatBN7.2 (Ensembl release 108) | 2022-11-10 |
| Rhodobacter sphaeroides | ASM1640v1 (Ensembl Bacteria release 55) | 2022-11-06 |
| Schizosaccharomyces pombe | ASM294v2 (PomBase update 2022-11-07) | 2022-11-08 |
| Xenopus tropicalis | UCB_Xtro_10.0 (Ensembl release 108) | 2022-11-05 |
All rRNAs and lncRNAs are retrieved from RNA central v21.
Standardized tRF ID of one tRF sequence is determined by tDRnamer. When one tRF sequence can be matched to multiple standardized IDs, only the ID in "tDR Name" column is used, and all IDs in "Synonyms" column are discarded.
All tRF-RNA interactions in tRFtarget are predicted using tRFtarget-pipeline v0.3.0.
Command options for calling tRFtarget-pipeline
--e_rnahybrid -15 --e_intarna -5 -b 5 -s 6
We add the manually curated gene-level experimental evidences to related binding sites predictions. For one tRF and one target gene in one gene-level evidence entry, we add this gene-level evidence to all predicted binding sites between all recorded tRFs with exactly the same sequence and all recorded transcripts of this specific target gene.
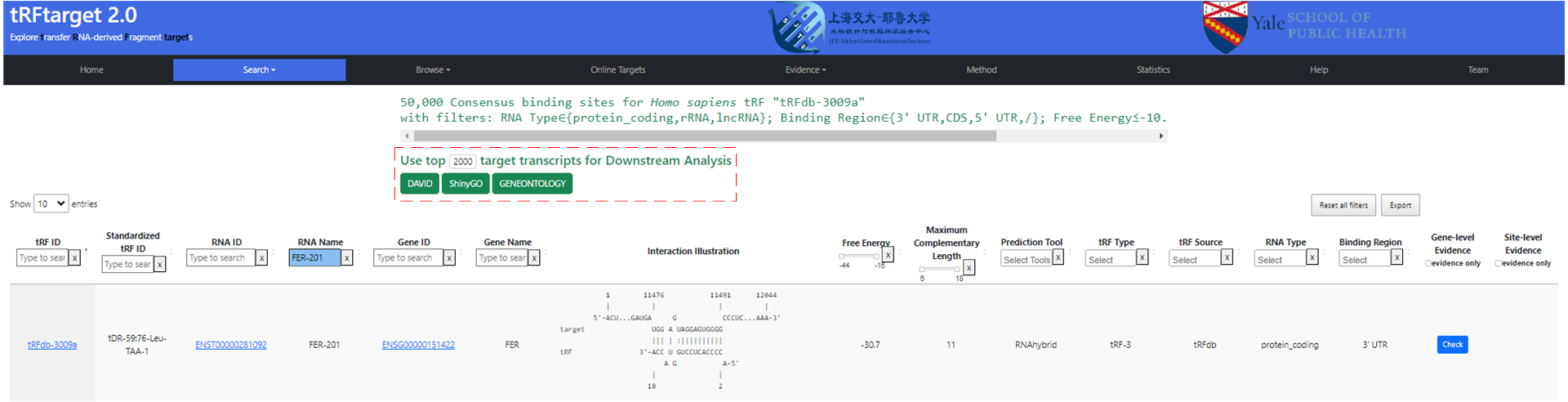
Users can effortlessly conduct downstream analysis with a single click after obtaining target genes for a specific tRF in one search (see figure below). The steps to select TOP genes for downstream analysis are: